Note
Go to the end to download the full example code.
Using a distance matrix¶
Generating data¶
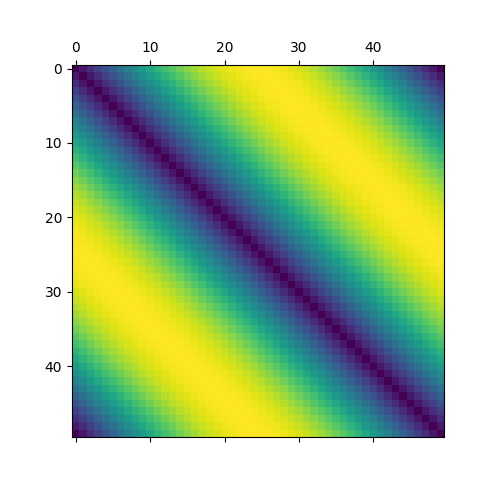
For this example we will generate evenly distributed points on a circle then compute the distance matrix of those points. In practice it makes little sense to go through this song and dance unless your data is given as a distance matrix.
Projecting our data¶
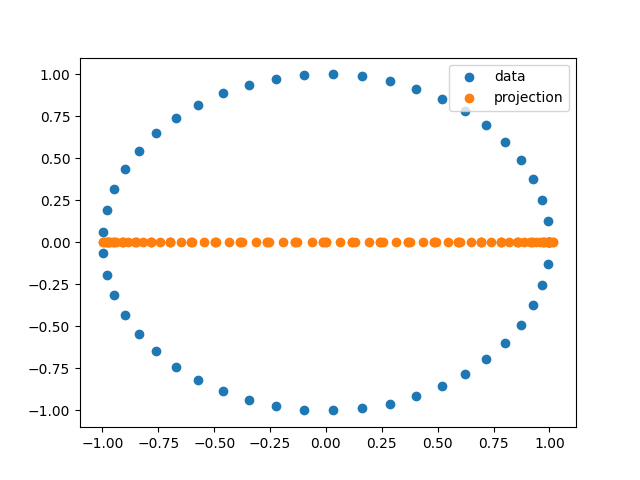
From here we will pretend that we don’t know the underlying data which generated the distance matrix. We pick [MDS](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.MDS.html) as our lense function as it plays nicely with distance matrices.
from sklearn.manifold import MDS
projection: np.ndarray = MDS(
n_components=1,
metric="precomputed",
n_init=1, # type: ignore
init="classical_mds",
).fit_transform(data)
plt.scatter(circle[:, 0], circle[:, 1], label="data")
plt.scatter(projection, np.zeros_like(projection), label="projection")
plt.legend()
plt.show()
Defining a clusterer¶
This is the last place where we need to do something different to facilitate running mapper on a distance matrix. All we have to do is tell the base clusterer that the metric is precomputed. From there the adapter should be clever enough to detect this fact.
from sklearn.cluster import AgglomerativeClustering
import zen_mapper as zm
sk = AgglomerativeClustering(
linkage="single",
n_clusters=None, # type: ignore
distance_threshold=0.2,
metric="precomputed",
)
clusterer = zm.sk_learn(sk)
Computing the mapper graph¶
cover_scheme = zm.Width_Balanced_Cover(n_elements=3, percent_overlap=0.4)
result = zm.mapper(
data=data,
projection=projection,
cover_scheme=cover_scheme,
clusterer=clusterer,
dim=1,
)
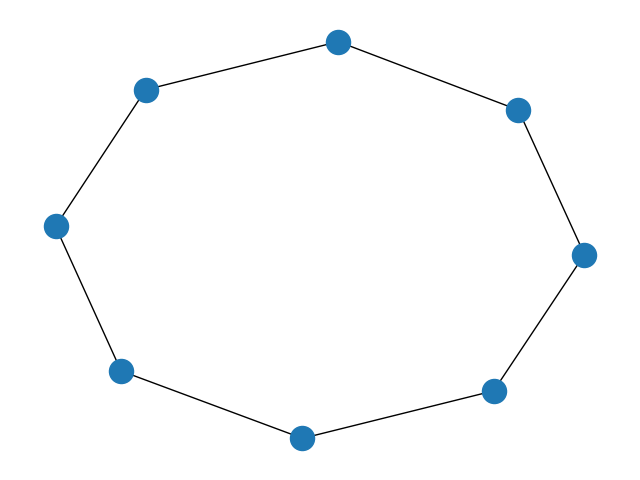
Visualizing the mapper graph¶
import networkx as nx
graph = zm.to_networkx(result.nerve)
nx.draw(graph)
Total running time of the script: (0 minutes 0.152 seconds)