Note
Go to the end to download the full example code.
Extracting Data from Nodes¶
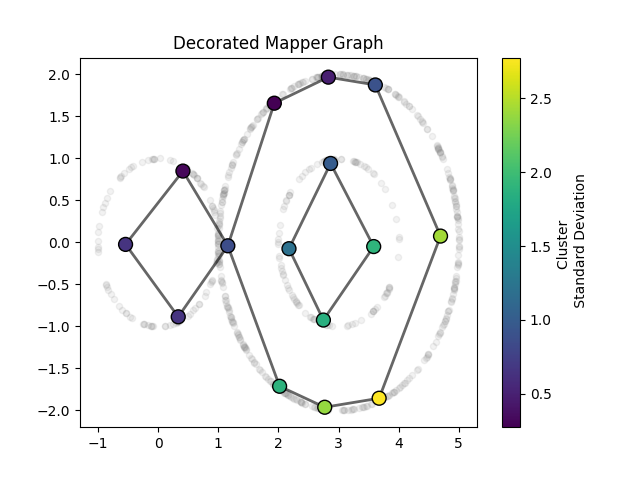
In this example we overlay a resulting mapper graph on top of the scatter-plot of datapoints. The nodes will be placed at the mean of the corresponding cluster. The color of the node will correspond to the standard deviation.
Generate Data¶
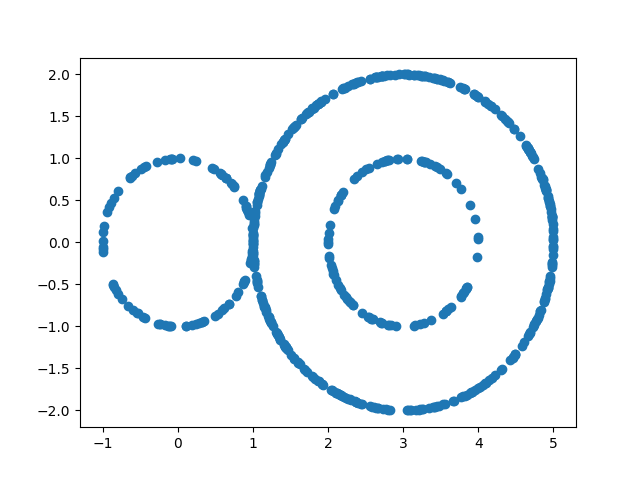
Here we will use sphere generator from kaiju mapper to generate a dataset
import matplotlib.pyplot as plt
import numpy as np
from kaiju_mapper.datasets import sphere
rng = np.random.default_rng(123)
circles = (
sphere(dim=1, num_samples=100, seed=rng),
sphere(dim=1, num_samples=400, radius=2, center=(3, 0), seed=rng),
sphere(dim=1, num_samples=100, radius=1, center=(3, 0), seed=rng),
)
data = np.vstack(circles)
plt.scatter(data[:, 0], data[:, 1])
plt.show()
Implement Mapper¶
from sklearn.cluster import DBSCAN
import zen_mapper as zm
cover_scheme = zm.Width_Balanced_Cover(n_elements=7, percent_overlap=0.2)
projection = data[:, 0]
clusterer = zm.sk_learn(DBSCAN(eps=0.5))
result = zm.mapper(
data=data,
projection=projection,
clusterer=clusterer,
cover_scheme=cover_scheme,
dim=1,
)
Accessing Data Points from Nodes¶
The MapperResult.nodes attribute holds a list of clusters which are
each a list of data indices belonging to the cluster.
Once you have access to these indices you can run your
own analysis.
cluster_centers = []
cluster_stds = []
# Get data from nodes!
for data_indices in result.nodes:
# This is where we extract data
cluster_points = data[data_indices]
center = np.mean(cluster_points, axis=0)
std = np.std(cluster_points)
cluster_centers.append(center)
cluster_stds.append(std)
node_positions = np.array(cluster_centers)
node_stds = np.array(cluster_stds)
Visualize¶
We can access all k-simplices by calling the MapperResult.nerve[k] attribute. This will return simplices stored as a (k+1)-tuple
of node_ids. One can then apply the same process as above to access the corresponding data points for analysis/visualization.
For now, we will just use this to draw edges manually.
plt.scatter(data[:, 0], data[:, 1], alpha=0.1, c="gray", s=20)
# draw edges from nerve attribute
for node1_id, node2_id in result.nerve[1]:
x_coords = [node_positions[node1_id, 0], node_positions[node2_id, 0]]
y_coords = [node_positions[node1_id, 1], node_positions[node2_id, 1]]
plt.plot(x_coords, y_coords, "k-", alpha=0.6, linewidth=2)
# draw nodes
scatter = plt.scatter(
node_positions[:, 0],
node_positions[:, 1],
c=node_stds,
s=100,
cmap="viridis",
edgecolors="black",
linewidth=1,
zorder=2, # just places nodes above edges
)
plt.colorbar(scatter, label="Cluster \n Standard Deviation")
plt.title("Decorated Mapper Graph")
plt.show()
Total running time of the script: (0 minutes 0.183 seconds)